ggpubr
data
> head(expr)
bcr_patient_barcode dataset GATA3 PTEN XBP1 ESR1 MUC1
1 BRCA1 BRCA 2.87 1.36 2.98 3.08 1.65
2 BRCA2 BRCA 2.17 0.43 2.55 2.39 3.08
3 BRCA3 BRCA 1.32 1.31 3.02 0.79 2.98
4 BRCA4 BRCA 1.84 0.81 3.13 2.50 -1.92
5 BRCA5 BRCA -6.03 0.25 -1.45 -4.86 -1.17
6 BRCA6 BRCA 1.80 1.31 4.04 2.80 3.53
基本格式
library(ggpubr)
# GATA3
ggboxplot(expr, ## data
x = "dataset",
y = "GATA3",
title = "GATA3", ## title
ylab = "Expression", ## ylab
color = "dataset", ## 配色分组
palette = "jco") ## 调色板,包括“grep”,brewer palattes,custom color palattes,ggsci
y = c("GATA3", "PTEN", "XBP1") # Create a list of plots
# Create a list of plots
p <- ggboxplot(expr, x = "dataset",
y = c("GATA3", "PTEN", "XBP1"),
title = c("GATA3", "PTEN", "XBP1"),
ylab = "Expression",
color = "dataset", palette = "jco")
# View GATA3
p$GATA3
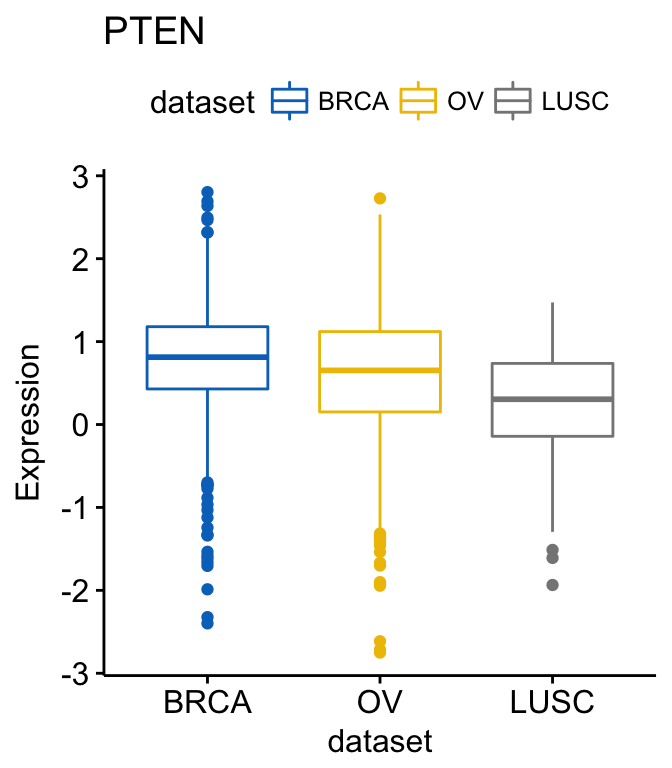
# View PTEN
p$PTEN
# View XBP1
p$XBP1
Add P-values anf Significance Levels
> list(c("BRCA", "OV"), c("OV", "LUSC"))
[[1]]
[1] "BRCA" "OV"
[[2]]
[1] "OV" "LUSC"
my_comparisons <- list(c("BRCA", "OV"), c("OV", "LUSC"))
p + stat_compare_means(comparisons = my_comparisons)
> combn(names(table(expr$dataset)),2,simplify = F)
[[1]]
[1] "BRCA" "LUSC"
[[2]]
[1] "BRCA" "OV"
[[3]]
[1] "LUSC" "OV"
> samples <- c("GATA3", "PTEN", "XBP1")
> combn(samples,2,simplify = F)
[[1]]
[1] "GATA3" "PTEN"
[[2]]
[1] "GATA3" "XBP1"
[[3]]
[1] "PTEN" "XBP1"
> combn(samples,2,simplify = T)
[,1] [,2] [,3]
[1,] "GATA3" "GATA3" "PTEN"
[2,] "PTEN" "XBP1" "XBP1"
For each of the genes, you can compare the different groups as follow:
> compare_means(c(GATA3, PTEN, XBP1) ~ dataset, data = expr)
# A tibble: 9 x 8
.y. group1 group2 p p.adj p.format p.signif method
<fct> <chr> <chr> <dbl> <dbl> <chr> <chr> <chr>
1 GATA3 BRCA OV 1.17e-177 3.50e-177 < 2e-16 **** Wilcoxon
2 GATA3 BRCA LUSC 6.80e- 73 1.40e- 72 < 2e-16 **** Wilcoxon
3 GATA3 OV LUSC 2.98e- 8 3.00e- 8 3.0e-08 **** Wilcoxon
4 PTEN BRCA OV 7.01e- 5 7.00e- 5 7.0e-05 **** Wilcoxon
5 PTEN BRCA LUSC 1.08e- 16 3.20e- 16 < 2e-16 **** Wilcoxon
6 PTEN OV LUSC 1.25e- 7 2.50e- 7 1.2e-07 **** Wilcoxon
7 XBP1 BRCA OV 2.49e-123 7.50e-123 < 2e-16 **** Wilcoxon
8 XBP1 BRCA LUSC 1.92e- 42 3.80e- 42 < 2e-16 **** Wilcoxon
9 XBP1 OV LUSC 4.49e- 11 4.50e- 11 4.5e-11 **** Wilcoxon
select or remove items (here cancer types)
item 指的是数据框的一行,这里对应x axis
select = c("BRCA", "OV")
remove = "BRCA"
指定items order
order = c("LUSC", "OV", "BRCA")
horizontal plots
rotate = TRUE


 浙公网安备 33010602011771号
浙公网安备 33010602011771号